Blog
Within BioPharm.Si programme, the project RDP1: Cell Line Quality Control for Biologics Production leaded by Prof Dr Kristina Gruden (National Institute of Biology, Department of Biotechnology and Systems Biology) aims to efficiently improve and/or select Chinese hamster ovary (CHO) cell lines with high production and long term stability. For parental CHO cell line improving, we used omics approach to identified genes that represent a bottleneck in the manufacturing process. The next step is to knockout these genes using CRISPR/Cas9.
CRISPR/Cas9 is a powerful-targeted genome editing technology that has revolutionised life science. During the last 5 years is apparently being used by everyone. Actually it has been applied in almost all fields of live science i.e basic research, biotechnology, and medicine (Fig 1) and thousands of papers using CRISPR/Cas9 have been published since 2011.
Fig 1. Applications of CRISPR/Cas9 (Hsu et al. Development and Applications of CRISPR-Cas9 for Genome Engineering, Cell, 2014)
But what is CRISP/Cas9 system? And what is it that turned CRISPR/Cas9 into the most efficient and highly used tool for targeted genome editing at a record-breaking speed?
Targeted genome editing technology allows inducing precise deletions, insertions or modifications of DNA sequences in a broad range of cell types and organisms. The first required step to apply the technology is to induce unique DNA double-strand breaks (DSBs) that will then be repaired by the machinery that almost all cell naturally possess. Depending on the repair pathway and the availability of a DNA donor template, we can accomplish different gene modifications. Two different DSB repair pathways have been described: non-homologous end joining which usually leads to gene knockout and homologous recombination that can be used for gene modification and gene insertion.
As we mentioned above, to apply genome engineering technology we first need to induce DSBs at the specific site of DNA that we want to edit. For this, synthetic nucleases such us zinc finger nucleases (ZFNs) and transcription activator-like effector nucleases (TALENs) were developed and have been available for already long time. These synthetic nucleases allow us to introduce DNA breaks in specific sites of the genome however they have not been widely accepted by the scientific community mostly because the design and construction of these enzymes is time consuming and expensive. It wasn’t until 2012, with the demonstration that CRISPR/Cas9 can be repurposed as a tool to generate programmable DSBs in DNA, that genome engineering technology became routine in molecular biology labs.
In fact CRISPR/Cas9 system was not “newly invented” by scientist but we borrowed it from bacteria. It’s a prokaryotic immune system that helps bacteria fight foreign genetic elements (e.g viral DNA). When a virus infect a bacterial cell for the first time the bacteria cuts the viral genome and incorporate a piece of it into its genome, at the CRISPR locus. When the CRISPR locus is transcribed, it produces the CRISPR targeting RNA (crRNA) that forms a complex with a Cas nuclease. The next time the bacterial cell will encounter the same virus, the crRNA will direct Cas nuclease to cut the foreign DNA. Nowadays we are using the same mechanism to generate specific breaks into the DNA of the organism we want to edit. To that, two main components have to be expressed in our target cells: the Cas9 nuclease and an engineered single guide RNA (sgRNA) that will guide Cas9 to the complementary target (Fig 2).
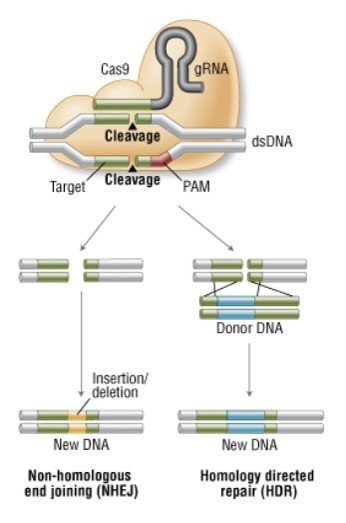
Fig 2. Genome engineering with CRISPR/Cas 9 system (New England BioLabs)
The simplicity is one of the main advantages of CRISPR/Cas9 technology. For each gene we want to edit, we only need to choose and design a sgRNA that will be co-expressed with the Cas9 nuclease. The relatively easy design and construction is even more evident when we compare it with other nucleases such us ZFNs and TALENs that require re-designing a new protein for each target gene. High efficiency and broad applicability are also advantages of the CRISPR/Cas9 that together with simplicity positioned this technology as a revolutionary tool for genome editing. On the other side, one of the few criticisms of the CRISPR/Cas9 system is the relatively high frequency of off-target mutations. However, a lot of research and advances in optimizing the system are going on in that direction.
Iskanje
Zadnje
- Workshop on "High Content Imaging and Data Science for Virtual Screening and Drug Discovery"
- VIS-NIR Low-cost Raman spectrometer
- Continuous bioprocess of bacteria and bacteriophages
- CRISPR/Cas9 tool for Genome Editing
- What researchers do at the Department of Catalysis and Chemical Reaction Engineering










